Researchers uncover how “silent” genetic changes drive cancer
At any given moment, the human genome spells out thousands of genetic words telling our cells which proteins to make. Each word is read by a molecule known as a tRNA.
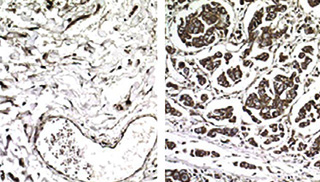
Cancer gone aggressive: In these samples from non-metastatic (left) and metastatic breast cancer (right), cells producing EXOSC2, a metastasis-promoting protein, are stained with a brown dye. The researchers found that EXOSC2 expression is enhanced in metastatic tumors because their cells have increased levels of a tRNA called GluUUC.
“We’ve long thought of these molecules as little more than middle men participating in the process of translating proteins,” explains Sohail Tavazoie, Leon Hess Associate Professor and head Rockefeller University’s Elizabeth and Vincent Meyer Laboratory of Systems Cancer Biology. But new research conducted in his lab suggests tRNA dynamics may play an important role in modulating the types of proteins in a cell.
In work published this week in Cell, Tavazoie and his colleagues describe how fluctuations in the amount of tRNAs can have a dramatic impact not only on the fate of a single cell, but also on diseases like metastatic breast cancer.
Spelling matters
Some words in the English language can be spelled in more than one way—like “color” and “colour,” which mean the same thing. The same is true for our genetic code. When cells activate a gene to make protein, their genome spells out three-letter words known as codons. Each codon is specific for one building block of a protein. But in some cases, two or several codons signify the same building block.
The codons are read by tRNAs, with one tRNA corresponding to each spelling. All living things use the same language, but codon usage and tRNA availability varies among different organisms and different tissues—just as the spelling “colour” predominates among the British, while “color” is more common among Americans.
Subtle changes have big consequences
For decades, researchers have assumed that these tRNAs had little impact on the function of the cell. But these molecules have been difficult to study, and Tavazoie’s team wanted to know if changes in the amount of tRNAs can affect cellular function.
“Traditionally, it has been hard to use standard methods to quantify the amount of tRNA in the cell,” says Tavazoie. The lead authors of the article, Hani Goodarzi, formerly a postdoc in the lab and now a new assistant professor at UCSF, and research assistant Hoang Nguyen, devised and applied a new method that utilizes state-of-the-art genomic sequencing technology to measure the amount of tRNAs in different cell types.
The team chose to compare breast tissue from healthy individuals with tumor samples taken from breast cancer patients—including both primary tumors that had not spread from the breast to other body sites, and highly aggressive, metastatic tumors.
They found that the levels of two specific tRNAs were significantly higher in metastatic cells and metastatic tumors than in primary tumors that did not metastasize or healthy samples. “There are six different ways to encode for the protein building block arginine,” explains Tavazoie. “Yet only one of those—the tRNA that recognizes the codon CGG—was associated with increased metastasis.”
The tRNA that recognizes the codon GAA and encodes for a building block known as glutamic acid was also elevated in metastatic samples.
The team hypothesized that the elevated levels of these tRNAs may in fact drive metastasis. Working in mouse models of primary, non-metastatic tumors, the researchers increased the production of the tRNAs, and found that these cells became much more invasive and metastatic. They also did the inverse experiment, with the anticipated results: reducing the levels of these tRNAs in metastatic cells decreased the incidence of metastases in the animals.
Driving metastasis
How do two tRNAs drive metastasis? The researchers teamed up with members of the Rockefeller University proteomics facility to see how protein expression changes in cells with elevated levels of these two tRNAs. “We found global increases in many dozens of genes,” says Tavazoie, “so we analyzed their sequences and found that the majority of them had significantly increased numbers of these two specific codons.”
According to the researchers, two genes stood out among the list. Known as EXOSC2 and GRIPAP1, these genes were strongly and directly induced by elevated levels of the specific glutamic acid tRNA. “When we mutated the GAA codons to GAG— a “silent” mutation because they both spell out the protein building block glutamic acid—we found that increasing the amount of tRNA no longer increased protein levels,” explains Tavazoie. These proteins were found to drive breast cancer metastasis.
The work challenges previous assumptions about how tRNAs function and suggests that tRNAs can modulate gene expression, according to the researchers. Tavazoie points out that “it is remarkable that within a single cell type, synonymous changes in genetic sequence can dramatically affect the levels of specific proteins, their transcripts, and the way a cell behaves.”
This research was supported by the National Institutes of Health.
 Cell, online: June 2, 2016 Cell, online: June 2, 2016Modulated Expression of Specific tRNAs Drives Gene Expression and Cancer Progression Hani Goodarzi, Hoang C.B. Nguyen, Steven Zhang, Brian D. Dill, Henrik Molina, Sohail F. Tavazoie |


