How CRISPR promotes antibiotic resistance in bacteria
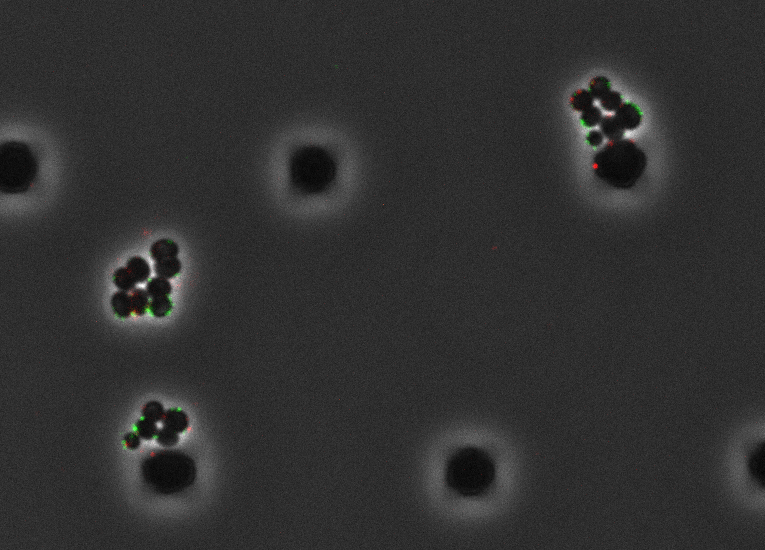
Bacteriophages (green, red) infecting Staphylococcus aureus cells (black clusters). Credit: Raphael Hofmann.
When Staphylococcus aureus bacteria are not causing pneumonia or idling harmlessly in our nostrils, they are busy fighting their own battles against a constant onslaught of phage virus bent on infecting them. An ancient adaptive immune system known as CRISPR, celebrated by scientists for its ability to cut and paste genes in the lab, is remarkable for its ability to record a memory of infection into the bacterium’s own DNA in order to slice the genomes of infecting viruses and eradicate them. CRISPR keeps most of the offending phage at bay.
Now, a new study in Nature highlights one of the consequences of CRISPR immunity. Every time a bacterium uses CRISPR, the researchers found, there’s a chance of mutations creeping into the bacterial genome itself. Some mutations are harmless; others kill the cell. But occasionally a fortuitous mutation can confer resistance to other phages, or even antibiotics.
“Whenever a cell uses CRISPR to defend itself, there’s some probability of it generating mutations that make it develop antibiotic resistance,” says Luciano Marraffini, Kayden Family Professor at Rockefeller and an investigator with the Howard Hughes Medical Institute. “It is exciting that we found this in a major human pathogen, Staphylococcus aureus, one known to develop antibiotic resistance. We can now explain why resistance may be happening.”
CRISPR is best known for its role in precision genome editing in the lab, but the CRISPR system that scientists use is in fact one of several CRISPR systems that help defend bacteria in nature. Not all of these systems are terribly precise. “Unlike the well-known type II CRISPR system, of which the Cas9 nuclease is used in genome editing, the type III CRISPR system doesn’t much care what DNA sequence it cuts,” says Charlie Mo, a postdoctoral fellow in Marraffini’s lab. Cells are left to repair the damage caused by their own immune systems.
Curious about how this non-specific system impacts the host bacteria, Marraffini and Mo exposed Staphylococcus aureus cells to phage and watched the type III system come online. As predicted, the cells’ DNA-repair systems tried to fix the damage caused by CRISPR’s non-specific DNA cutting and occasionally failed. “No system is foolproof,” Marraffini says. “Every time you cleave DNA, there’s a chance of introducing mutations during repair.”
Mutations are one of the principal drivers of evolution in bacteria. But what may be good news for the evolution of Staphylococcus aureus is bad news for us. Constantly mutating bacteria tend to stumble upon antibiotic resistance sooner or later. It is plausible, Marraffini says, that mutations caused by type III CRISPR are among the major factors in the transformation of harmless Staphylococcus aureus bacteria into more deadly antibiotic-resistant variants.
This would be particularly interesting, given that CRISPR immune systems prevent another major driver of bacterial evolution—horizontal gene transfer, or the exchange of large chunks of genetic material between cells. Phages that infect multiple bacteria can ferry beneficial genes between hosts, but only if they manage to do so before they are eradicated by CRISPR. Marraffini suspects that the mutations introduced by DNA repair offset that loss—ensuring that, although the opportunity for horizontal gene transfer via phage is lost, bacteria still have a chance to mutate and benefit from further evolution.
For Marraffini and Mo, the next step is figuring out whether the type III system is driving mutations—and antibiotic resistance—in other human pathogens. “Mycobacterium tuberculosis, the bacterium that causes tuberculosis, is another pathogen that carries type III CRISPR,” Marraffini says. “Future studies will address whether other organisms with type III CRISPR demonstrate similarly elevated levels of mutations that can lead to antibiotic resistance.”



