Vertebrate Genomes Project Plan
The VGP will be completed based on taxonomic hierarchy, which is a relative ranking of a group of organisms beginning with the largest classification, domain, to the smallest classification, species: orders (Phase 1), families (Phase 2), and genera (Phase 3) to eventually all species (Phase 4). This strategy will allow for genomic analyses to be performed at increasing levels of phylogenetic scale. Phase 1 of the VGP will focus on the generation of high-quality near error-free genomes of 260 species representing all vertebrate orders with a divergence time of ~50 million years ago (MYA) or greater from their most recent common ordinal ancestor, including human and some species on the brink of extinction.
The VGP Project Phases
- All orders (phase 1, ~260 species);
- All families (phase 2, ~1,000 species);
- All genera (phase 3, ~10,000 species); and
- All species (phase 4, ~66,000 species).
In Phase 1, we will select one representative species from each order, which amounts to a total of ~260 vertebrate species, many of which are endangered. We will sequence the heterogametic sex (when it exists) so that both sex chromosomes can be assembled for each species. Species selection will be based on a combination of criteria, including species with existing draft genomes in need of improvement, equal representation of taxa based on systematic classification, even coverage of divergence times, and prominent use in biomedical research.
The VGP will be complete in these four specific project phases to allow us to gain scientific insight at each phase and to continue to integrate emerging technologies. Additionally, we expect our approaches and questions at each phase to lead to the development of new algorithms, including algorithms for genome assemblies, alignments, annotations, comparative genomics, etc., which would then be applied to the next phase. Lastly, this approach will help us secure the needed funds in stages through grants and other fundraising efforts. The success of Phase 1 will allow us to leverage the necessary funds to sequence the genomes of phases 2, 3, and 4, all ~1K vertebrate families, then all ~10K genera, and finally all ~66K species, respectively.
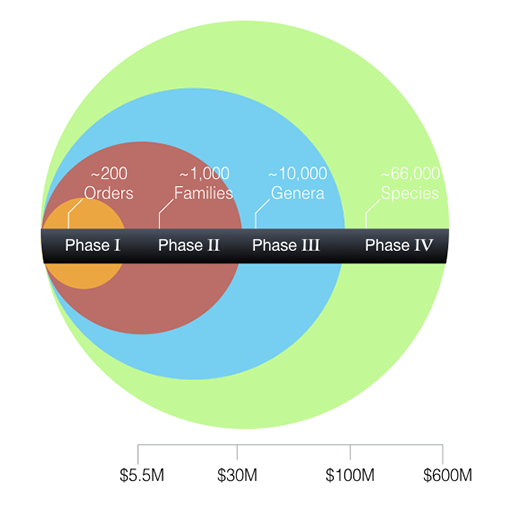
Vertebrate phylogeny classification numbers and the planned phases of the VGP.
Once funding is secured for all ~260 species in Phase 1, we will be able to generate approximately 12 genomes per week equating to 6-8 months of sequencing and assembly. Collection of the high quality tissue samples not yet identified will occur in parallel, adding another 4-6 months of time. Alignments and annotations will occur in parallel, and at 10 genomes per week, will add another 6 months of time. Together, we expect to complete all 260 species plus the 4 invertebrate
outgroups within 1.5 years from the start of a major source of funding. Biological analyses for publications will occur simultaneously, although some analyses can only occur after annotation and alignment of the 260 species, which will add another 12 months before submitting papers for publication.
- Over 260 near-gapless, chromosome-level and phased genome assemblies representing all extant vertebrate orders;
- New computational and bioinformatic resources that the entire scientific community can use;
- Major scientific discoveries in phylogenomics, evolution, comparative genomics, genetics of specialized traits, and biomedicine; and
- Publications in special issues of high- and intermediate-profile journals that reach a wide audience and bring greater public awareness to the benefits of science;
We will also complete various broad-strokes analyses of evolutionary differences in genes. For example, we will generate the first ordinal-level genome-scale phylogeny tree across vertebrates. We will also be able to make predictions about which species are most at risk for extinction.
All genome data generated by the VGP will be made publicly available in Genome Ark and through existing databases (agreements made with NCBI, ENSEMBL, and UCSC) on the Amazon and Microsoft cloud, with our informatics and data management platform hosted by DNA Nexus.



